Tracking a virus villain
Federal COVID-19 funding allocated by the North Carolina General Assembly is supporting vital research by Carolina scientists, who are keeping tabs on the deadly coronavirus using wastewater and nasal test swabs.
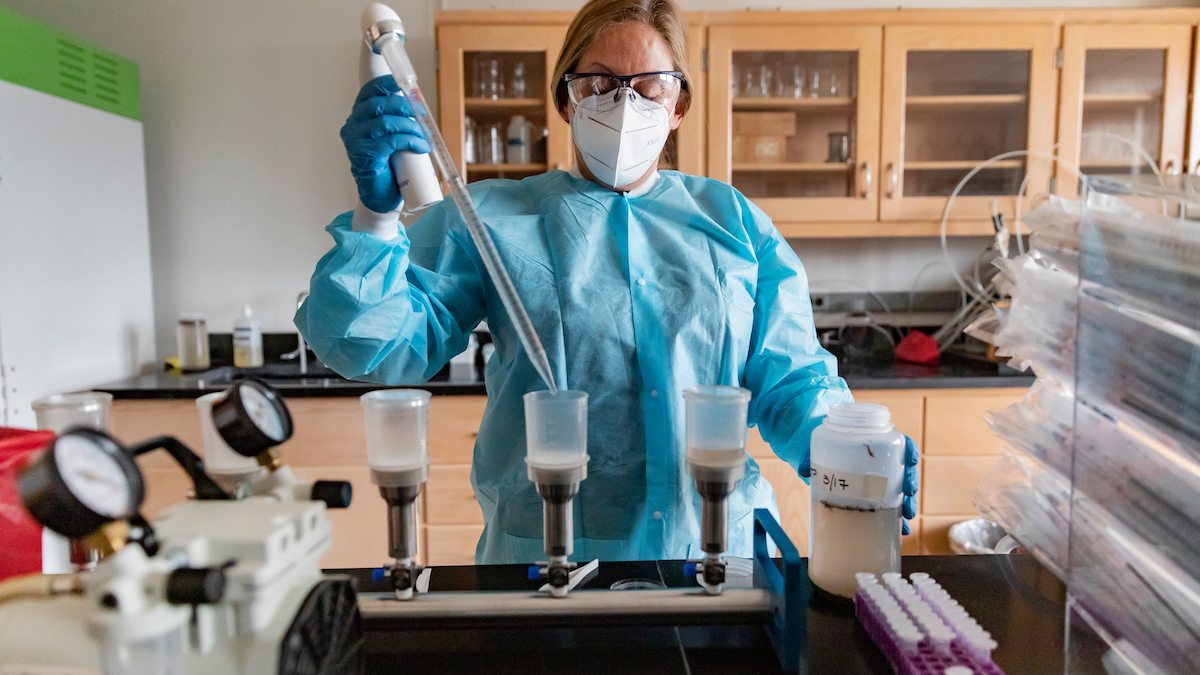

COVID-19 is a tricky, slippery villain. Sometimes it looks like a bad cold or the flu; at other times, a heart attack or stroke. All over the world, it eludes the heroes in the white coats by changing its identity, location by location.
Clearly, this viral villain requires vigilant surveillance. And the implementation of “alternative COVID-19 surveillance methods” is just what the North Carolina General Assembly singled out when it allocated $15 million of the latest federal COVID-19 funding to the North Carolina Policy Collaboratory based at Carolina. The $15 million allocation approved by the state legislature in March is subject to Centers for Disease Control and Prevention approval since the funding is allocated by the CDC Epidemiology and Laboratory Capacity cooperative funding agreement to the North Carolina Department of Health and Human Services to detect, respond to, control and prevent infectious diseases.
“DHHS needs to ensure that a statewide surveillance effort is statistically representative of the North Carolina population as a whole in terms of demographics and geography,” said Jeffrey D. Warren, executive director of the Collaboratory. “This is one reason that the Collaboratory is proposing a partnership between the School of Medicine and UNC Health to collect data because its network has providers in half of all North Carolina ZIP codes.”
The North Carolina General Assembly established the Collaboratory in 2016 to put the research expertise across the UNC System to practical use by state and local governments. When COVID-19 became a concern in 2020, the legislature appropriated $29 million to the Collaboratory to support a wide range of research projects on treatment, community testing and prevention of COVID-19.
“Some of the new funding will go to determining not only the best way to track these variants but also the broadest way to do that, to generate data from across the state of North Carolina and not just at a few key locations,” said Rachel Noble of the Institute of Marine Sciences and the Gillings School of Global Public Health. “We’re out to serve more of North Carolina.”
Two Carolina researchers whose projects received funding in the first round in 2020 are hoping to expand their efforts with the new funding focused on surveillance of SARS-CoV-2 — the virus that causes COVID-19. Noble of the and Dirk Dittmer of the School of Medicine are already keeping tabs on the virus and its variants using wastewater and nasal test swabs, respectively, with plans to expand their dragnets.
And in keeping with the Collaboratory’s mission, they will be putting this knowledge to timely, practical use.
“We’re not out to conduct epidemiological research as part of this new funding. This new funding is for implementing surveillance and monitoring technologies,” said Noble, who has a joint appointment as Mary and Watts Hill Jr. Distinguished Professor at the Institute of Marine Sciences, the Institute for the Environment and the Gillings School of Global Public Health.
Surveillance by sequencing
Dittmer is using the test swabs to sequence the genomes of the virus strains circulating in the state to make sure that the tests being used will be able to identify the emerging variants. He aims to have a system in place before the state’s schools and colleges begin their fall sessions.

Dirk Dittmer
“The Collaboratory is working with the North Carolina Department of Health and Human Services, the UNC System and other stakeholders to develop and implement a fast, local and comprehensive ‘surveillance-by-sequencing’ approach that will deliver a representative picture of which SARS-CoV-2 viral variants are circulating in our state and how they are distributed across geographic regions and with regard to risk groups of our population on a weekly basis,” said Dittmer, professor of microbiology and immunology and director of viral genomics at the School of Medicine. Dittmer is also a member of the UNC Lineberger Comprehensive Cancer Center.
It’s important to be able to identify the genetics of different strains of the virus because some strains may be more easily spread than others and cause more disease, leading to longer stays in the hospital and a need for more intensive care units. Since the beginning of the epidemic, Carolina has contributed more than half of all SARS-CoV-2 sequences for the state of North Carolina.
Dittmer’s lab has also collected blood serum from COVID-19 patients and compared it to virus tests on the same patients to see if patients with antibodies to the virus can still get sick and infect others. This information is helpful for future vaccine development.
Wastewater tracking
Professor Rachel Noble works in her environmental microbiology lab at the Institute of Marine Sciences in Morehead City on March 30, 2021. Noble is the director of the IMS Morehead City field site and a Mary and Watts Hill, Jr. Distinguished Professor.
(Johnny Andrews/UNC-Chapel Hill)
Noble has worked with a team of researchers to build the capacity for wastewater-based surveillance across the state. Wastewater-based surveillance allows the team to look at the trends of infection across whole communities rather than at the individual clinical sample level. The funding from 2020 allowed her team to develop rapid response methods for tracking SARS-CoV-2 in wastewater from over 20 locations throughout the state.
In 2021, Noble’s lab is now adding to the analysis by using wastewater to spot new strains (variants) in a way that complements Dittmer’s research. Samples collected from municipal wastewater systems capture virus variants more broadly since people who are asymptomatic and wouldn’t seek out testing (common in the younger age groups) are all making “contributions” to her research.
“If you’ve flushed a toilet, then you’ve contributed,” she explained.
Noble’s goal is to formalize this surveillance in areas across the state so that local governments can use the findings to make decisions about opening up their communities and so health care systems can prepare for new virus variations and outbreaks. Aggregate testing from wastewater analysis can also provide a continuous supply of positive viral samples for genomic sequencing, critical for identifying current and emerging variants.
Noble has been working with researchers at other UNC System schools — including Appalachian State, NC State University, East Carolina University, UNC Charlotte and UNC Wilmington — to collect and analyze the wastewater in their areas.
“So the first thing that any additional funds would allow us to do would be to ramp up the surveillance in municipalities and in counties where wastewater surveillance data is not yet available,” Noble said. “The second goal is to get a handle on the evolution of variants or mutations of the SARS-CoV-2 virus across North Carolina.”
Noble is partnering with diagnostic testing companies to test for different variants. “These groups are designing tests quickly as variants of concern emerge, so the turnaround on the diagnostic capability is amazing,” she said. And Noble’s lab has the wastewater samples that can show them how effective the tests are.
For example, when the British variant emerged, Noble contacted a biotechnology company to ask if they had designed a test for it.
“We actually just designed them yesterday,” they told her.
“Great, could you send us some for free?” she asked them. “And we’ll test them out, and we’ll tell you how they worked.”
“Things are happening so fast. That’s really how it happened,” Noble recalled. “We have a very small lab, so we don’t have time to be miring through thousands — millions, actually — of RNA sequences of clinical samples to design diagnostic tests. And so they’re very interested in serving a role in helping us to understand the details of the variants.”
North Carolina is one of eight states currently feeding wastewater surveillance information to the CDC, which would like to eventually establish a national wastewater surveillance system, Noble said. While the focus is on COVID-19 for the moment, a national system would have much broader implications.
“The CDC is really thinking about how to use wastewater surveillance more broadly in the future — maybe to detect a hepatitis outbreak or to track antibiotic resistance, for example, in a major metropolitan city, or to better target education campaigns,” she said.
But Noble is focused on what her research can do for the state now. “Some of the new funding will go to determining not only the best way to track these variants but also the broadest way to do that, to generate data from across the state of North Carolina and not just at a few key locations. We’re out to serve more of North Carolina.”




